Push model using huggingface_hub.
Browse files
README.md
CHANGED
|
@@ -12,7 +12,7 @@ tags:
|
|
| 12 |
---
|
| 13 |
|
| 14 |
# ibm/biomed.sm.mv-te-84m-MoleculeNet-ligand_scaffold-CLINTOX-101
|
| 15 |
-
**
|
| 16 |
|
| 17 |
- **Developers:** IBM Research
|
| 18 |
- **GitHub Repository:** [https://github.com/BiomedSciAI/biomed-multi-view](https://github.com/BiomedSciAI/biomed-multi-view)
|
|
@@ -22,9 +22,8 @@ tags:
|
|
| 22 |
|
| 23 |
## Model Description
|
| 24 |
|
| 25 |
-
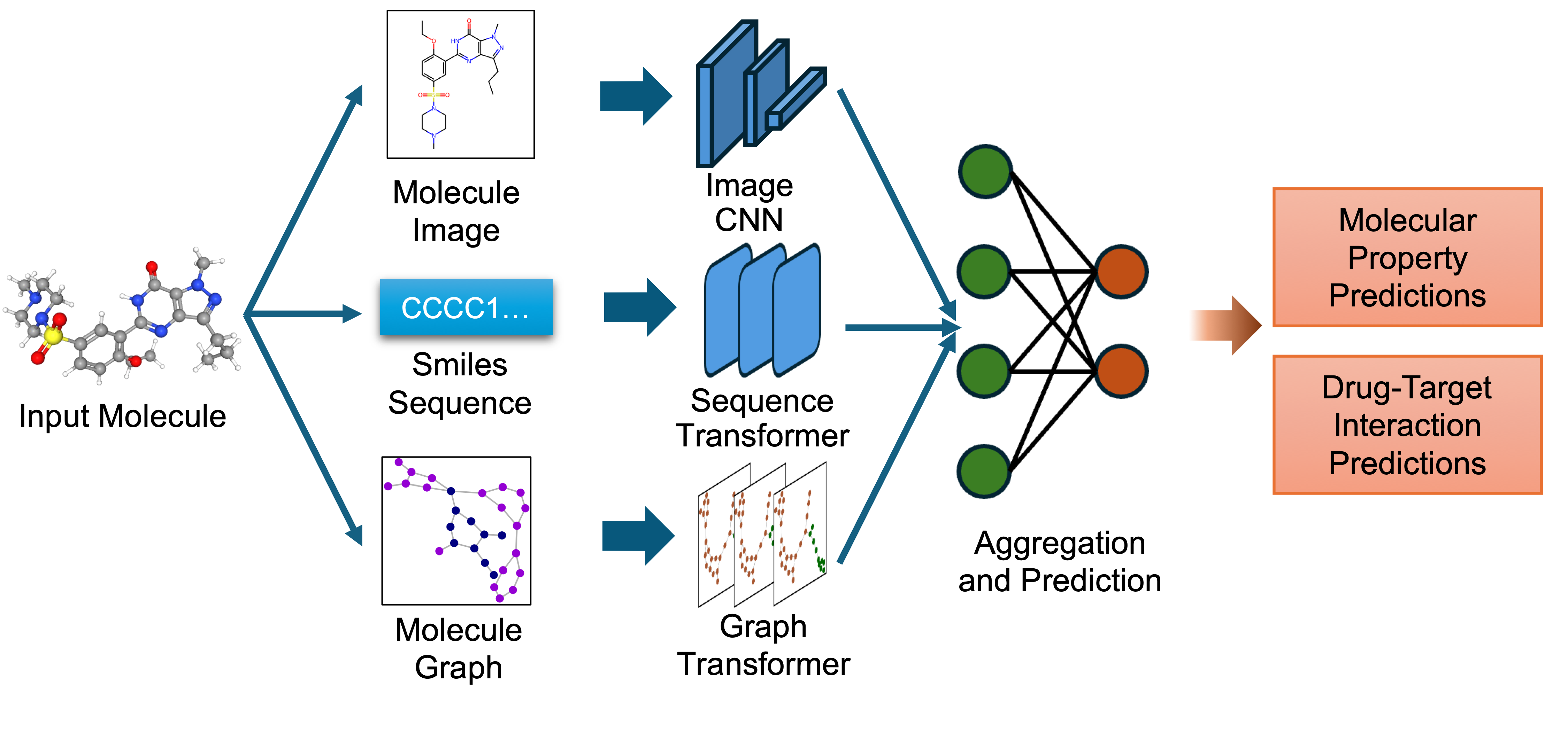
`biomed.sm.mv-te-84m` is a biomedical foundation model for small molecules created using MMELON (Multi-view Molecular Embedding with Late Fusion), a flexible approach to aggregate multiple views (sequence, image, graph) of molecules in a foundation model setting. While models based on single view representation typically performs well on some downstream tasks and not others, the multi-view model performs robustly across a wide range of property prediction tasks encompassing ligand-protein binding, molecular solubility, metabolism and toxicity. It has been applied to screen compounds against a large (> 100 targets) set of G Protein-Coupled receptors (GPCRs) to identify strong binders for 33 targets related to Alzheimer’s disease, which are validated through structure-based modeling and identification of key binding motifs [Multi-view biomedical foundation models for molecule-target and property prediction](https://arxiv.org/abs/2410.19704).
|
| 26 |
|
| 27 |
-
Source code is made available in [this repository](https://github.com/BiomedSciAI/biomed-multi-view).
|
| 28 |
|
| 29 |
|
| 30 |
|
|
@@ -39,7 +38,7 @@ The embeddings from these single-view pre-trained encoders are combined using an
|
|
| 39 |
The model is intended for (1) Molecular property prediction. The pre-trained model may be fine-tuned for both regression and classification tasks. Examples include but are not limited to binding affinity, solubility and toxicity. (2) Pre-trained model embeddings may be used as the basis for similarity measures to search a chemical library. (3) Small molecule embeddings provided by the model may be combined with protein embeddings to fine-tune on tasks that utilize both small molecule and protein representation. (4) Select task-specific fine-tuned models are given as examples. Through listed activities, model may aid in aspects of the molecular discovery such as lead finding or optimization.
|
| 40 |
|
| 41 |
|
| 42 |
-
The model’s domain of applicability is small, drug-like molecules. It intended for use with molecules less than 1000 Da molecular weight. The MMELON approach itself may be extended to include proteins and other macromolecules but does not at present provide embeddings for such entities. The model is at present not intended for molecular generation. Molecules must be given as a valid SMILES string that represents a valid chemically bonded graph. Invalid inputs will impact performance or lead to error.
|
| 43 |
|
| 44 |
## Usage
|
| 45 |
|
|
|
|
| 12 |
---
|
| 13 |
|
| 14 |
# ibm/biomed.sm.mv-te-84m-MoleculeNet-ligand_scaffold-CLINTOX-101
|
| 15 |
+
`biomed.sm.mv-te-84m` is a biomedical foundation model for small molecules created using **MMELON** (**M**ulti-view **M**olecular **E**mbedding with **L**ate Fusi**on**), a flexible approach to aggregate multiple views (sequence, image, graph) of molecules in a foundation model setting. While models based on single view representation typically performs well on some downstream tasks and not others, the multi-view model performs robustly across a wide range of property prediction tasks encompassing ligand-protein binding, molecular solubility, metabolism and toxicity. It has been applied to screen compounds against a large (> 100 targets) set of G Protein-Coupled receptors (GPCRs) to identify strong binders for 33 targets related to Alzheimer’s disease, which are validated through structure-based modeling and identification of key binding motifs [Multi-view biomedical foundation models for molecule-target and property prediction](https://arxiv.org/abs/2410.19704).
|
| 16 |
|
| 17 |
- **Developers:** IBM Research
|
| 18 |
- **GitHub Repository:** [https://github.com/BiomedSciAI/biomed-multi-view](https://github.com/BiomedSciAI/biomed-multi-view)
|
|
|
|
| 22 |
|
| 23 |
## Model Description
|
| 24 |
|
|
|
|
| 25 |
|
| 26 |
+
Source code for the model and finetuning is made available in [this repository](https://github.com/BiomedSciAI/biomed-multi-view).
|
| 27 |
|
| 28 |

|
| 29 |
|
|
|
|
| 38 |
The model is intended for (1) Molecular property prediction. The pre-trained model may be fine-tuned for both regression and classification tasks. Examples include but are not limited to binding affinity, solubility and toxicity. (2) Pre-trained model embeddings may be used as the basis for similarity measures to search a chemical library. (3) Small molecule embeddings provided by the model may be combined with protein embeddings to fine-tune on tasks that utilize both small molecule and protein representation. (4) Select task-specific fine-tuned models are given as examples. Through listed activities, model may aid in aspects of the molecular discovery such as lead finding or optimization.
|
| 39 |
|
| 40 |
|
| 41 |
+
The model’s domain of applicability is small, drug-like molecules. It is intended for use with molecules less than 1000 Da molecular weight. The MMELON approach itself may be extended to include proteins and other macromolecules but does not at present provide embeddings for such entities. The model is at present not intended for molecular generation. Molecules must be given as a valid SMILES string that represents a valid chemically bonded graph. Invalid inputs will impact performance or lead to error.
|
| 42 |
|
| 43 |
## Usage
|
| 44 |
|